eDNA METABARCODING OF STOMACHS FROM TWO CANADIAN CLEANER FISHES: WILD CUNNER WRASSES AND LUMPFISH FROM ATLANTIC SALMON SEACAGES
Lumpfish (Cyclopterus lumpus) and cunner wrasse (Tautogolabrus adspersus) are two species of cleaner fish used in sea cages by Canadian Atlantic salmon aquaculture for the biological control of the ectoparasitic salmon louse (Lepeophtheirus salmonis). Lumpfish are the predominant species used in Canadian salmon aquaculture, whereas cunner are the only member of the cleaner wrasse family that inhabit Atlantic waters. To better characterize the diets of wild cunner populations and cultured lumpfish from industrial-scale sea cages (lumpfish) we combined traditional morphological diet assessments with eDNA metabarcoding. The primary goal of this research was to provide insight into how natural diet variation and prey availability influence the propensity of cleaner fish to feed on salmon lice in an industrial aquaculture setting.
We first performed traditional morphological stomach analysis following eDNA protocols and then isolated DNA from 122 lumpfish stomachs from Canadian Atlantic salmon cages (Cooke Aquaculture Inc.) in southern Newfoundland1 and 110 cunner gastrointestinal tracts from four regions along coastal Atlantic Canada2. Diet assessments and taxonomic identifications were completed using morphological techniques (biomass; prey abundance), and further characterized by molecular metabarcoding for targeted identification at higher taxonomic resolutions (read abundance: READS; sequence diversity: ESVs).
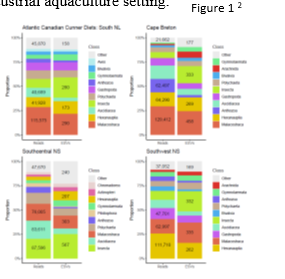
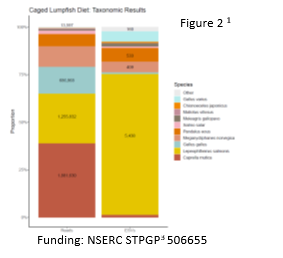
Wild cunners, fed primarily on benthic prey including invasive ascidians (southern regions), barnacles, bivalves, and malacostracans (Figure 1). Lumpfish diets were largely characterized by pelagic wild crustaceans, and industrial feed components (fishmeal and poultry), indicating they fed predominately in the water column (Figure 2). Most notably, metabarcoding was the only technique able to identify the salmon louse in ethanol-preserved lumpfish or cunner stomachs, and document when cleaning activity had occurred. For both cleaners, traditional diet assessments produced low resolution, semi-quantitative prey abundance at the order or family level, whereas metabarcoding could identify prey that were present at the species level. This analysis indicated a clear difference in diet between the two species which can be used to make decisions on their use within Canadian sea cages.